- ImageCLEF 2025
- LifeCLEF 2025
- ImageCLEF 2024
- LifeCLEF 2024
- ImageCLEF 2023
- LifeCLEF 2023
- ImageCLEF 2022
- LifeCLEF2022
- ImageCLEF 2021
- LifeCLEF 2021
- ImageCLEF 2020
- LifeCLEF 2020
- ImageCLEF 2019
- LifeCLEF 2019
- ImageCLEF 2018
- LifeCLEF 2018
- ImageCLEF 2017
- LifeCLEF2017
- ImageCLEF 2016
- LifeCLEF 2016
- ImageCLEF 2015
- LifeCLEF 2015
- ImageCLEF 2014
- LifeCLEF 2014
- ImageCLEF 2013
- ImageCLEF 2012
- ImageCLEF 2011
- ImageCLEF 2010
- ImageCLEF 2009
- ImageCLEF 2008
- ImageCLEF 2007
- ImageCLEF 2006
- ImageCLEF 2005
- ImageCLEF 2004
- ImageCLEF 2003
- Publications
- Old resources
You are here
ImageCLEFmed: Medical classification
Medical compound figure separation and multi-label classification task
The objective of this task is to work on compound figures of the biomedical literature and to separate them if possible and/or attach to the sub parts labels about the content.
Schedule
- 12.11.2014: registration opens for all ImageCLEF tasks (until 30.04.2015)
- 15.03.2015: data release
- 01.05.2015: deadline for submission of runs by the participants
- 15.05.2015: release of processed results by the task organizers
- 07.06.2015: deadline for submission of working notes papers by the participants
- 30.06.2015: notification of acceptance of the working notes papers
- 15.07.2015: camera ready working notes papers
- 08.-11.09.2015: CLEF 2015, Toulouse, France
Citations
- When referring to ImageCLEFmed 2015 task general goals, general results, etc. please cite the following publication:
- Alba García Seco de Herrera, Henning Müller and Stefano Bromuri , Overview of the ImageCLEF 2015 medical classification task, in: CLEF working notes 2015, Toulouse, France, 2015.
-
BibText:
@InProceedings{GMB2015,-
Title = {Overview of the {ImageCLEF} 2015 medical classification task},
Author = {Garc\'ia Seco de Herrera, Alba and M\"uller, Henning and Bromuri, Stefano},
Booktitle = {Working Notes of {CLEF} 2015 (Cross Language Evaluation Forum)},
Year = {2015},
Month = {September},
Location = {Toulouse, France}}
- Alba García Seco de Herrera, Henning Müller and Stefano Bromuri , Overview of the ImageCLEF 2015 medical classification task, in: CLEF working notes 2015, Toulouse, France, 2015.
- When referring to ImageCLEFmed task in general, please cite the following publication:
- Jayashree Kalpathy-Cramer, Alba García Seco de Herrera, Dina Demner-Fushman, Sameer Antani, Steven Bedrick and Henning Müller, Evaluating Performance of Biomedical Image Retrieval Systems –an Overview of the Medical Image Retrieval task at ImageCLEF 2004-2014 (2014), in: Computerized Medical Imaging and Graphics
-
BibText:
@Article{KGD2014,-
Title = {Evaluating Performance of Biomedical Image Retrieval Systems-- an Overview of the Medical Image Retrieval task at {ImageCLEF} 2004--2014},
Author = {Kalpathy--Cramer, Jayashree and Garc\'ia Seco de Herrera, Alba and Demner--Fushman, Dina and Antani, Sameer and Bedrick, Steven and M\"uller, Henning},
Journal = {Computerized Medical Imaging and Graphics},
Year = {2014}}
Motivation
An estimated 40% of the figures in PubMed Central are compound figures (images consisting of several sub figures) like the images above. When data of articles is made available digitally, often the compound images are not separated but made available in a single block. Information retrieval systems for images should be capable of distinguishing the parts of compound figures that are relevant to a given query. A major step for making the content of the compound figures accessible is the detection of compound figures and then their separation into sub figures that can subsequently be classified into modalities and made available for research.
The medical classification task of ImageCLEF 2015 uses a subset of PubMed Central.
Task overview
There are four types of tasks in 2015:
- Compound figure detection:
Compound figure identification is therefore a required first step to make available compound images from the literature. Therefore, the goal of this task is to identify whether a figure is a compound figure or not. The task makes training data available containing compound and non compound figures from the biomedical literature. - Multi-label classification:
Characterization of compound figures is difficult, as they may contain subfigures from various imaging modalities or image types. This task aims to label each compound figure with each of the modalities (of the 30 classes of a defined hierarchy shown below) of the subfigures contained without knowing where the separation lines are. - Figure separation:
This task was first introduced in 2013. The task makes available training data with separation labels of the figures and then a test data set where the labels will be made available after the submission of the results. In 2015, a larger number of compound figures is distributed compared to the previous task. - Subfigure classification:
Similar to the modality classification task organized in 2011-2013 this task aims to classify images into the 30 classes of the hierarchy shown below. The images are the subfigures extracted from the compound figures distributed for the figure separation task.
Classification hierarchy
The following hierarchy is used for the modality classification, same classes as in ImageCLEF 2012-2013, although this year '[COMP] Compound or multiplane images' is not a class.
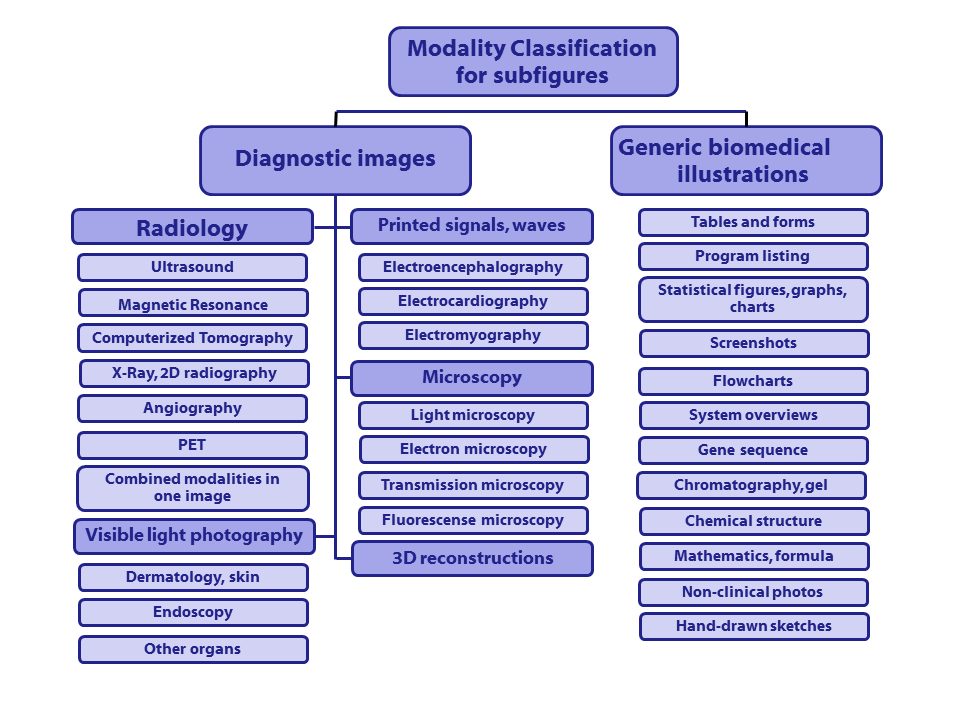
Class codes with descriptions (class codes need to be specified in run files):
([Class code] Description)
- [Dxxx] Diagnostic images:
- [DRxx] Radiology (7 categories):
- [DRUS] Ultrasound
- [DRMR] Magnetic Resonance
- [DRCT] Computerized Tomography
- [DRXR] X-Ray, 2D Radiography
- [DRAN] Angiography
- [DRPE] PET
- [DRCO] Combined modalities in one image
- [DVxx] Visible light photography (3 categories):
- [DVDM] Dermatology, skin
- [DVEN] Endoscopy
- [DVOR] Other organs
- [DSxx] Printed signals, waves (3 categories):
- [DSEE] Electroencephalography
- [DSEC] Electrocardiography
- [DSEM] Electromyography
- [DMxx] Microscopy (4 categories):
- [DMLI] Light microscopy
- [DMEL] Electron microscopy
- [DMTR] Transmission microscopy
- [DMFL] Fluorescence microscopy
- [D3DR] 3D reconstructions (1 category)
- [Gxxx] Generic biomedical illustrations (12 categories):
- [GTAB] Tables and forms
- [GPLI] Program listing
- [GFIG] Statistical figures, graphs, charts
- [GSCR] Screenshots
- [GFLO] Flowcharts
- [GSYS] System overviews
- [GGEN] Gene sequence
- [GGEL] Chromatography, Gel
- [GCHE] Chemical structure
- [GMAT] Mathematics, formulae
- [GNCP] Non-clinical photos
- [GHDR] Hand-drawn sketches
Data collection
The dataset used in this task is a subset of images contained in articles from the biomedical literature extracted from the PubMed Central.
- Compound figure detection:
20,000 images labelled as compound figures or not compound figures are distributed. - Figure separation:
A subset of the compound figures from the compound figure detection task is distributed to be separated into subfigures. - Multi-label classification:
A subset of the compound figures are distributed for the multi-label task. - Subfigure classification:
Figures from the multi-label classification task are separated into subfigures and each of the subfigures are labelled.
If the figure ID is "1297-9686-42-10-3", then the corresponding subfigures IDs are "1297-9686-42-10-3-1", "1297-9686-42-10-3-2", "1297-9686-42-10-3-3" and "1297-9686-42-10-3-4"
Submission instructions
Compound figure detection
The format of the result submission for the Compound figure detection subtask should be the following:
1471-2458-10-S1-S4-3 COMP 0.9
1471-2458-10-52-5 COMP 1
1471-2458-11-133-1 COMP 0.4
1423-0127-17-34-8 NOCOMP 0.8
1465-9921-6-21-6 NOCOMP 0.9
...
where:
- The first column contains the figure-ID (IRI). This ID does not contain the file format ending and it should not represent a file path.
- The second column is the classcode.
- The third column represents the normalized score (between 0 and 1) that your system assigned to the specific result.
You need to respect the following constraints:
- Each specified figure has to be part of the collection (dataset).
- A figure cannot be contained more than once.
- All figures of the test set have to be contained in the runfile.
- Only known class codes are accepted.
Please note that each group is allowed to submit a maximum of 10 runs.
Compound figure separation
The format of the result submission for the compound figure separation subtask has to be an XML file with the following structure :
where:
- The root element is <annotations>.
- The root contains one <annotation> element per image. Each one of these elements has to contain :
- A <filename> element with the name of the compound image (IRI) (excluding the file extension)
- One or more <object> elements that define the bounding box of each subfigure in the image. Each <object> must contain :
- 4 <point> elements that define the 4 corners of the bounding box. The <point> elements must have two attributes (x and y), which correspond to the horizontal and vertical pixel position, respectively. The preferred order of the points is :
- top-left
- top-right
- bottom-left
- bottom-right
You also need to respect the following constraints:
- Each specified image has to be part of the collection (dataset).
- An Image cannot appear more than once in a single XML results file.
- All the images of the testset must be contained in the runfile.
- The resulting XML file MUST validate against the XSD schema that will be provided.
Multi-label classification
The format of the result submission for the Multi-label classification classification subtask should be the following:
1751-0147-52-24-3 DRMR DRXR
1475-925X-6-10-8 DRUS GHDR
1471-2210-10-7-3 GCHE
1475-2875-6-10-2 GFIG
...
where:
- The first column contains the figure-ID (IRI). This ID does not contain the file format ending and it should not represent a file path.
- The rest of the columns contain the classcode.
You need to respect the following constraints:
- Each specified figure has to be part of the collection (dataset).
- A figure cannot be contained more than once.
- All figures of the test set have to be contained in runfile.
- Only known classcodes are accepted.
Please note that each group is allowed a maximum of 10 runs.
Subfigure classification
Similar to the compound figure detection subtask, the format of the result submission for the Subfigure classification subtask should be the following:
1743-422X-4-12-1-4 D3DR 0.9
1471-2156-8-36-3-4 GFIG 1
1475-2859-9-86-6-1 GFIG 0.4
1475-2840-10-59-2-4 DMLI 0.8
...
where:
- The first column contains the subfigure-ID (IRI). This ID does not contain the file format ending and it should not represent a file path.
- The second column is the classcode.
- The third column represents the normalized score (between 0 and 1) that your system assigned to the specific result.
You need to respect the following constraints:
- Each specified subfigure has to be part of the collection (dataset).
- A subfigure cannot be contained more than once.
- All subfigures of the test set have to be contained in runfile.
- Only known class codes are accepted.
Please note that each group is allowed a maximum of 10 runs.
Evaluation methodology
Compound Figure Separation : The Java Archive (JAR) containing the application to run the evaluation can be downloaded below. The source code of the application is also contained in the ZIP archive.
Download Compound Figure Separation Evaluation Tool & Source
More details on the evaluation will be provided at a later time
Results
| Compound figure detection |
| Group name | Run | Run type | Correctly classified in % |
| Biomedical Computer Science Group | task1_run2_mixed_sparse1 | mixed | 85.39 |
| Biomedical Computer Science Group | task1_run1_mixed_stemDict | mixed | 83.88 |
| Biomedical Computer Science Group | task1_run3_mixed_sparse2 | mixed | 80.07 |
| Biomedical Computer Science Group | task1_run4_mixed_bestComb | mixed | 78.32 |
| Biomedical Computer Science Group | task1_run6_textual_sparseDict | textual | 78.34 |
| CIS UDEL | exp1 | visual | 82.82 |
| Biomedical Computer Science Group | task1_run5_visual_sparseSift | visual | 72.51 |
| Multi-label classification |
| Group name | Run | Hamming Loss | |
| IIS | output_6 | 0.0817 | |
| IIS | output_8 | 0.0785 | |
| IIS | output_9 | 0.0710 | |
| IIS | output_7 | 0.0700 | |
| IIS | output_10 | 0.0696 | |
| IIS | output_5 | 0.0680 | |
| IIS | output_1 | 0.0678 | |
| IIS | output_3 | 0.0675 | |
| MindLAB | predictions_Mindlab_ImageclefMed_multilabel_test_comb2lbl | 0.0674 | |
| IIS | output_4 | 0.0674 | |
| IIS | output_2 | 0.0671 | |
| MindLAB | predictions_Mindlab_ImageclefMed_multilabel_test_comb1lbl | 0.0500 |
| Figure separation |
| Group name | Run | Run type | Correctly classified in % |
| NLM | run2_whole | visual | 84.64 |
| NLM | run1_whole | visual | 79.85 |
| AAUITEC | aauitec_figsep_combined | visual | 49.40 |
| AAUITEC | aauitec_figsep_edge | visual | 35.48 |
| AAUITEC | aauitec_figsep_bond | visual | 30.22 |
| Subfigure classification |
| Group name | Run | Run type | Correctly classified in % |
| Biomedical Computer Science Group | task4_run5_train_20152013.txt | mixed | 67.60 |
| Biomedical Computer Science Group | task4_run4_clean_rf.txt | mixed | 67.24 |
| Biomedical Computer Science Group | task4_run1_combination.txt | mixed | 66.48 |
| Biomedical Computer Science Group | task4_run8_clean_short_rf.txt | mixed | 66.44 |
| Biomedical Computer Science Group | task4_run7_clean_comb_librf.txt | mixed | 65.99 |
| Biomedical Computer Science Group | task4_run6_clean_libnorm.txt | mixed | 64.34 |
| Biomedical Computer Science Group | task4_run3_textual.txt | textual | 60.91 |
| Biomedical Computer Science Group | task4_run2_visual.txt | visual | 60.91 |
| CMTECH | resultsSubfigureRunWholeCov.txt | visual | 52.98 |
| CMTECH | resultsSubfigure.txt | visual | 48.61 |
| BMET | sf_run_3.txt | visual | 45.63 |
| BMET | sf_run_6.txt | visual | 45.00 |
| BMET | sf_run_4.txt | visual | 44.34 |
| BMET | sf_run_2.txt | visual | 43.62 |
| BMET | sf_run_1.txt | visual | 37.56 |
| BMET | sf_run_5.txt | visual | 37.56 |
Organizers
- Alba García Seco de Herrera, National Library of Medicine (NLM/NIH), Bethesda, MD, USA, albagarcia(replace-by-an-at)nih.gov
- Stefano Bromuri, University of Applied Sciences Western Switzerland in Sierre, Switzerland, stefano.bromuri(replace-by-an-at)hevs.ch
Acknowledgements
| Attachment | Size |
|---|---|
| 278.73 KB |






